
"Noisy" data can be identified by the presence of multiple peaks and numerous "N"s within your sequence. You may need to reprep your sample to sufficiently remove one or more inhibitory components to obtain any sequence data. Please check the Contaminant chart in the Template preparation and purification section for a list of potential inhibitors and the amounts that are tolerable. Solutions: The cycle sequencing reaction used to amplify samples for automated sequencing is very sensitive to the presence of certain contaminants, some of which will completely inhibit our sequencing enzyme.

While our sequencers are very sensitive and can detect a range of DNA concentrations, there is still a "threshold" amount that must be reached to obtain any sequence data. Check our What kinds of DNA can we sequence and how much do we need? section to make sure you’ve provided the appropriate amount of DNA and/or primer.
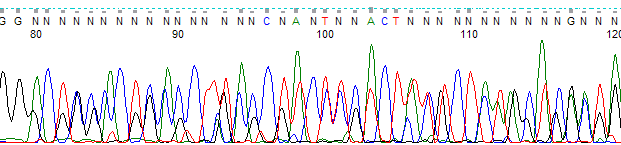
Solutions: Doublecheck your quantitations, stock concentrations and dilutions. Avoid areas where the peaks are broader and not well separated - this will occur towards the end of the sequence where the fragments are larger and the polymer cannot adequately resolve single nucleotides, causing inaccurate basecalling.Ĭause: Not enough or no DNA/primer in tube If you’ve designed your own custom primer from previous sequence data, make sure you were using a reliable area of sequence - look for sharp, well-defined peaks with no ambiguity. GL primer 1 can be used with the pGL2 vector but not the pG元 vector). T7, M13-48R), others are specific to one particular type (e.g. While many of the primers we provide are quite common to many different vectors (e.g. Solutions: if you’ve chosen one of the sequencing facility’s vector primers, make sure it is present in your vector. If it appears that you have done everything correctly and followed our suggestions, then look below for some additional reasons why you might obtain less than optimal DNA sequence data quality. If you are having trouble getting good sequencing results for your samples, you may first want to look through our Sequencing Basics section for some recommendations on template preparation and quantitation. The two most common causes for failure to get good or any sequence data for your samples are purity and concentration of your template DNA.
